Scientists have learned how to store images in DNA and then perfectly retrieve them
Apr 8, 2016
Share:

As our data creation and acquisition requirements soar, so must the technology to store it all, and a team of computer scientists and electrical engineers from the University of Washington have seem to have made a bit of a breakthrough.
In a paper presented in April at the ACM International Conference on Architectural Support for Programming Languages an Operating Systems (ASPLOS), the team detailed one of the first complete systems to encode, store and retrieve digital data using DNA molecules, capable of storing information millions of times more compactly than existing storage technologies.
In the paper, one outlined experiment details successfully encoding digital data from four image files into the nucleotide sequences of synthetic DNA snippets.
Even more significantly, they were also able to reverse the process and retrieve the correct sequences from a larger pool of DNA, reconstructing the original images without losing a single byte of information.
Life has produced this fantastic molecule called DNA that efficiently stores all kinds of information about your genes and how a living system works — it’s very, very compact and very durable.
We’re essentially repurposing it to store digital data — pictures, videos, documents — in a manageable way for hundreds or thousands of years.
– Luis Ceze, UW associate professor of computer science and engineering.
The first step of the process is converting all the ones and zeroes that make up the image using the four basic building blocks of DNA – adenine, guanine, cytosine and thymine. It sounds simple, but the issue has been working out how to pack as much data as they can into as short a string as possible while remaining error free.
This is done using simple Huffman coding, a common approach for lossless data compression.
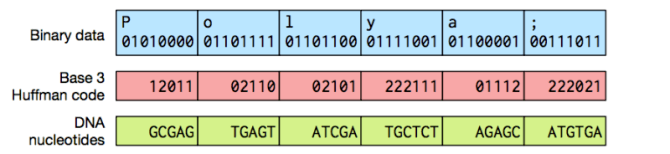
Once the team have determined how to best represent a file, artificial DNA is then synthesized based on their calculations. Getting the data back out is a much simpler process, due to distinctive markers placed within the DNA by the team denoting the start and end points of files, and just reverse the encoding process to reproduce the original file.
Information can be stored many millions of times more densely in DNA molecules than in any existing data storage technologies. Other technologies also degrade after just a few years or decades, whereas DNA can be dehydrated and reliably preserved for centuries.
It’s this long term reliability which makes this technology more suited to long term archival storage rather than the needs of general computer users where data needs to be accessed randomly and immediately.
It’s also somewhat expensive, and it’s not the kind of thing you can at home with a junior science kit. If the price can be brought down to something more manageable, however, it could help to solve the world’s impending data storage problem.
The digital universe is doubling in size every two years and will multiply 10-fold between 2013 and 2020 – from 4.4 trillion gigabytes to 44 trillion gigabytes.
Authors of the paper are UW computer science and engineering doctoral student James Bornholt, UW bioengineering doctoral student Randolph Lopez, UW associate professor of computer science and engineering Luis Ceze, UW associate professor of electrical engineering and of computer science and engineering Georg Seelig, and Microsoft researchers and UW CSE affiliate faculty Doug Carmean and Karin Strauss.
You can read the full paper, here.
[University of Washington via Gizmodo]
John Aldred
John Aldred is a photographer with over 25 years of experience in the portrait and commercial worlds. He is based in Scotland and has been an early adopter – and occasional beta tester – of almost every digital imaging technology in that time. As well as his creative visual work, John uses 3D printing, electronics and programming to create his own photography and filmmaking tools and consults for a number of brands across the industry.

































Join the Discussion
DIYP Comment Policy
Be nice, be on-topic, no personal information or flames.
3 responses to “Scientists have learned how to store images in DNA and then perfectly retrieve them”
I wanted to write a little comment to support you.
(Y)
One wonders whether our cat videos and instagram brunch shots really need to be preserved…